 Hi-C data allows examining the genome 3D organization at multiple scales (Rocha et al. 
As such in this review on big-data challenges in epigenomics, we will focus especially on datasets obtained from mammalian genomes, as well as on data analysis solutions used in this context. This results in very large and complex datasets, especially for large genomes, as the number of possible pairwise interactions increases exponentially with the genome length. Hi-C is the most comprehensive and high-throughput derivative, allowing us to score contact frequency between virtually any pair of genomic loci (Lieberman-Aiden et al. 2012), whereas 5C allows probing multiple pairwise interactions between predesigned anchor points (Phillips-Cremins et al. Among them, 4C allows detecting pairwise interactions between one target anchor point and potentially any other genomic region (van de Werken et al. 2006) or high-throughput sequencing have been proposed subsequently. However, other high-throughput derivatives of 3C based on microarrays hybridization (Dostie et al. This can be assessed with PCR, using a pair of primers specifically designed to target predefined regions, as per the original 3C protocol. By analyzing the resulting hybrid molecules, we can assess the physical interaction between distant genomic loci (Belton et al. The fact that they are ligated together at the end of the process indicates some degree of physical proximity at the beginning of the experimental procedure. The loose DNA fragment ends are then re-ligated, so as to obtain hybrid molecules, which may contain two fragments of DNA that were not adjacent but indeed far apart in the original linear genomic sequence. The technique is based on cross-linking of DNA and associated proteins to stabilize chromatin 3D structure, then digesting DNA with restriction enzymes. 3C allows probing physical interaction between non-adjacent genomic loci. 2002) and its high-throughput derivatives have been the most prominent ones. Among the latter, chromosome conformation capture (3C) (Dekker et al. 2017) and other molecular biology techniques. Use other annotations for more in-depth exploration of the results.Our knowledge of chromatin 3D organization has greatly increased over the past 20 years thanks to the development of novel experimental techniques, including high-resolution and high-throughput imaging techniques (Huang et al. 
You can zoom in or scroll around in the heatmap to better understand where the DIRs occur in relation to each other and on the chromatin map. The blue boxes on the heatmap represent the significant DIRs. Here we can see a screenshot of Juicebox with the Rao 2017 data loaded and our annotations visualized on the heatmap. The DIRs will appear as blue rectangles on the heatmap. Select chromosome 16 in both drop-down menus below the “Chromosomes” selection toolbar and click the refresh button. After loading it, the “rao2017Annotations.txt” annotations will be shown in the “Select 2D annotation file(s) to open” panel - select it and click the Open button to load in into Juicebox. tab/button, and navigate to the “rao2017Annotations.txt” file. Alternatively, use the View > Show Annotations Panel menu. This file has been pre-generated, right-click and save it from here.Īssuming the “Rao et al. | Cell 2017” “Auxin treated (6 hrs)” data is loaded, click the Show Annotation Panel button at the lower right corner of Juicebox window. This function will create a text file in your working directory containing the annotations which can then be imported into Juicebox. exportJuicebox(rao2017, logfc_cutoff = 2, logcpm_cutoff = 1, p.adj_cutoff = 0.001, file_name = "rao2017Annotations.txt") Say your Hicexp object is named rao2017 simply run the following line of code in R. 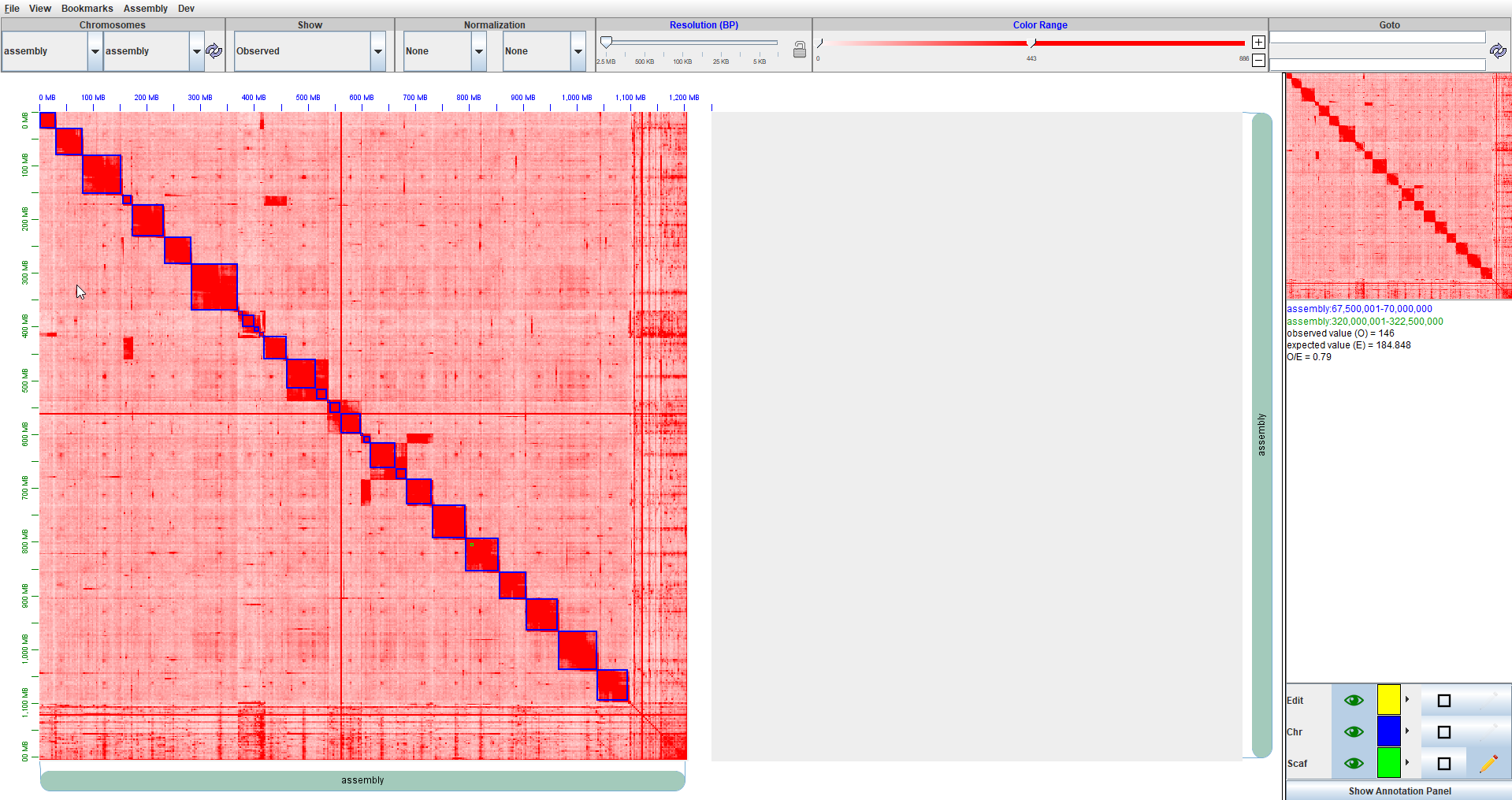
You will want to select the main map for “Auxin treated (6 hrs)”.Īssuming you have already analyzed the data using multiHiCcompare and you now have a Hicexp object containing the comparison results you can use the exportJuicebox function to export the list of DIRs as an annotations file for input into Juicebox. 
The Rao 2017 data are available in Juicebox by default under File > Open menu, “Rao et al. | Cell 2017”. We compared the data between the normal HCT-116 cells and the cells treated with auxin for 6 hours. For our example, we will use the data from Rao 2017. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
